The way our faces look depends on our sex, age, genetic inheritance and ethnic background. In more subtle ways, it depends on our nutritional status, psychological state and smoking and drinking habits. Lack of sleep and our general health, as we know too well, also affect how we look.
More fundamentally, facial appearance is subject to the effects of our genetic make-up, and when genes important in craniofacial development are altered in some way, we may be born with facial features that are different from typically developing children. Alcohol, anti-convulsive drugs and other teratogens are also known to induce facial differences in the developing foetus.

Discover B2B Marketing That Performs
Combine business intelligence and editorial excellence to reach engaged professionals across 36 leading media platforms.
The human brain possesses a remarkable ability to recognise what is familiar or unfamiliar in a face. The general public can recognise the face shape of individuals with Down’s syndrome and experienced clinical geneticists develop this innate skill to such a degree that they recognise subtle facial features associated with several hundred dysmorphic syndromes.
Many of these conditions are so rare that lack of exposure to clinical cases during training makes it hard for less experienced
clinicians to develop the skill of recognising the so-called facial gestalt that suggests the initial diagnosis and determines the examinations and tests needed to confirm that first hunch.
NEW TECHNIQUES
Novel 3D image analysis techniques can now visualise and recognise dysmorphic facial features, and they show considerable potential for training clinical geneticists and assisting the diagnosis of difficult cases, or in situations where local expertise is limited. They can also be useful in multi-disciplinary studies identifying the roles of individual genes and the pathological mechanisms of genetic conditions.
CT and MR images of the head can be used to reconstruct the shape of a face, but in many clinical situations, such modalities are inappropriate because of the unnecessary radiation exposure they involve or lack of patient cooperation arising from the underlying condition. In contrast, 3D laser and photogrammetric images of the face can be captured in a fraction of a second, involve negligible risk and often require only moderate cooperation.
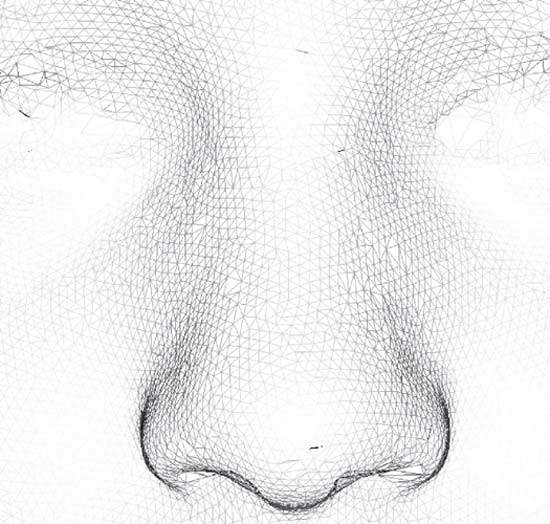
Such a 3D face image is typically made up of a mesh of tens or even hundreds of thousands of 3D points (Figure 1) representing the surface of the face and one or more high-resolution 2D colour images capturing facial appearance. In appropriate 3D viewing software, the face can be inspected from any direction, at any desired magnification and at a proximity that may be impossible or uncomfortable in the live clinical situation.
Craniofacial surgeons and dentists already use 3D photographs to monitor changes in face shape arising from surgery and orthodontic treatment. Whether it is required for corrective treatment or to assist diagnosis, it is often necessary to compare a child’s face with a group of faces of individuals of the same gender and of a similar age and ethnic background. By adding anatomical landmarks to a 3D image, anthropometric measurements can be derived and used to compare an individual’s face to
a set of appropriate norms.
Such measurements, though, are a considerable simplification of the shape complexity of a human face. Surface-based representations employing 25,000 or more 3D points capture much more subtle features. Of course, this potentially gives rise to a data analysis problem when several hundred faces are involved in the comparison.
However, using statistical data compaction techniques, such as principal component analysis (PCA), it is possible to build a model of the variation in face shape in a group of individuals that requires only 100 or so data values (called principal components or modes) to reconstruct each face surface with a high degree of accuracy. Then, individual or group comparisons of face shape only require an analysis of the differences in these 100 or so mode values.
Typically, the most significant mode of a PCA model of face shape corresponds to facial growth, and it can be used as a surrogate measure of age or of facial development. Other modes may capture overall shape differences in terms of the face being round or elliptical, with eyes close together or far apart, and with ears that are set low or high on the side of the head. Some genetic conditions, such as Noonan, Williams and 22q11 deletion syndromes not only involve face shape differences but also heart anomalies that require early intervention by a surgeon.
In Williams syndrome, there is a fullness around the eyes; the face is narrower, especially at the temples; the nose is shorter and more bulbous at the tip; the philtrum (central region between nostrils and upper lip) is longer; the lips are fuller; and the lower jaw is shorter.
In Noonan syndrome, the upper eyelids are ptotic (droopy); the eyes are more widely spaced and typically have lower outer corners; the bridge of the nose is flatter and the nose itself is somewhat broader; the lower jaw is smaller; and the ears are usually set lower on the head.
In the case of 22q11 deletion syndrome, surgical repair of an associated cleft palate may itself alter subsequent facial growth, resulting in a flatter profile to the mid-face.
It is important, therefore, to be aware that medical intervention can add to face shape variation arising from the effects of age, sex, altered genes, familial likeness, ethnicity and, of course, facial expression.
FACE CLASSIFICATION
An intuitive approach to classifying a face is to compute the average face shape of groups of children with different genetic conditions and then determine which average face a patient’s face is most similar to. This and other more mathematically sophisticated approaches are proving remarkably accurate at differentiating between children with some genetic conditions and controls with no underlying genetic anomaly.
Once these face shape models, known as dense surface models, have been shown to be effective at classifying face shape, they can be used to screen the faces of children for whom a diagnosis has not been made.
Of course, recognition of face shape does not imply a diagnosis. A diagnosis is made by an appropriately trained clinician backed up, whenever possible, by genetic testing. For some dysmorphic syndromes there is no definitive genetic test and a clinical diagnosis has to suffice. For others, for example Noonan syndrome, a number of important genes may have been identified, but mutations for those genes may not be found in some children for whom there is a compelling clinical diagnosis.
In such situations, confirmation of associated facial characteristics using the analysis of a 3D image will strengthen the clinical diagnosis and may even encourage a clinician to undertake the remaining tests when an incomplete subset of those available has thus far proven negative.
The discriminating accuracy of dense surface models of face shape is obviously dependent on the number and homogeneity of the 3D images collected. For example, too few images will result in an average face that is less representative of the variation in that particular syndrome and will reduce the model’s ability to classify faces correctly. The probability of correctly differentiating between the faces of affected and unaffected children using dense surface models is between 0.85 and 0.95
for the six or so conditions studied so far. As more images are collected, the classification accuracy typically increases.
The rarity of most of these genetic conditions makes it unacceptable simply to sit back and wait for affected children to turn up at a genetics or paediatric clinic. So it is necessary to travel extensively to meetings where large numbers of families with affected children gather to learn directly from each other or physicians about health, educational and social issues affecting their children.
Some clinical geneticists and paediatricians specialise in particular conditions and are able to call upon families, who often give generously of their time, to contribute to research projects. With this approach to patient recruitment, it is important to have a device that is relatively easy to transport by train or air. Moreover, it must be sufficiently robust to survive baggage handlers and airport security checks.
RESEARCH TOOL
Besides showing potential for assisting training in facial dysmorphology and diagnosing genetic conditions, 3D face shape analysis can also be of benefit when used in conjunction with behavioural and molecular analyses aimed at identifying the underlying genetic anomalies or pathobiological mechanisms of a genetic condition.
Within the phenotype of a genetic condition, there can be a wide range of effects in terms of severity or number of symptomatic features. Some children may have a severe form of a heart condition that is typical for a syndrome, while others may have milder or no cardiac features. Some children may have mild facial features compared with others, and there can be a wide range of mild to severe cognitive and behavioural effects.
It can be helpful in these situations to focus on children who
have a confirmed condition but who are in some way atypical: in their particular gene mutation, in the size of their deletion or in the way they are affected. In a research study of Williams syndrome, for example, it was possible to combine detailed facial, behavioural and molecular analyses of an atypical child to provide additional evidence suggesting that a particular gene is the likely cause of the face shape differences in the syndrome.
FUTURE DEVELOPMENTS
It is technically possible to reconstruct 3D face shape from 2D images, but obviously with some loss of 3D information. This may be an essential pre-requisite of facial analysis until 3D cameras and scanners are sufficiently cheap and generally available.
So, in the not too distant future, a clinician having difficulty with a genetic diagnosis may be able to email a 2D image to a website, where it will be converted to a 3D and compared with 3D models of facial dysmorphology for a large numbers of genetic conditions. A replying email might contain a list of potential diagnoses ordered according to similarity of facial morphology. This could inform subsequent clinical examinations and genetic testing, possibly even reducing the number, and
hence cost, of the tests undertaken.
Such a useful facility may itself be overtaken by current developments in micro-array technology that support the detailed examination of the entire genome of an individual on a single chip. Even so, there will still be important research to be undertaken in understanding craniofacial development and associated disease to which 3D facial analysis will continue to make a useful contribution for many years to come.





